## # A tibble: 100 × 20
## V1 V2 V3 V4 V5 V6 V7 V8 V9 V10
## <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 0.274 -0.0367 0.855 0.968 1.42 0.523 0.563 1.11 1.64 0.619
## 2 2.24 -0.546 0.234 -1.34 1.31 0.521 -0.610 -0.861 -0.270 0.230
## 3 -0.125 -0.607 -0.854 -0.149 -0.665 0.607 0.162 -0.863 0.604 1.19
## 4 -0.544 1.11 -0.104 1.02 0.700 1.66 0.490 0.0234 0.256 -0.127
## 5 -1.46 -0.274 0.112 -0.852 0.315 1.05 1.39 0.285 1.14 2.68
## 6 1.06 -0.754 -1.38 1.08 0.370 1.50 -0.360 -0.214 1.83 0.871
## 7 0.116 -0.966 0.274 0.0185 -0.210 0.544 -2.51 2.20 0.593 -1.09
## 8 0.390 0.400 -0.467 0.478 0.848 0.257 -0.0995 0.999 -0.193 -0.404
## 9 0.193 0.197 -1.24 1.91 0.115 0.452 -0.267 -0.273 -1.61 -0.326
## 10 -0.0964 1.18 2.55 0.956 0.0256 -0.0456 -0.659 0.0392 0.172 0.643
## # … with 90 more rows, and 10 more variables: V11 <dbl>, V12 <dbl>, V13 <dbl>,
## # V14 <dbl>, V15 <dbl>, V16 <dbl>, V17 <dbl>, V18 <dbl>, V19 <dbl>, V20 <dbl>
## # A tibble: 100 × 1
## V1
## <dbl>
## 1 -1.27
## 2 1.84
## 3 0.459
## 4 0.564
## 5 1.87
## 6 0.528
## 7 2.43
## 8 -0.895
## 9 -0.206
## 10 3.11
## # … with 90 more rows
## Perform ridge regression with 10-fold CV
ridge1_cv <- cv.glmnet(x = x_cont, y = y_cont, type.measure = "mse",
lambda = exp(seq(-5,-1.5, length.out = 100)),
nfold = 10, alpha = 0) # alpha is the e-net mixing par
## Penalty vs CV MSE plot
plot(ridge1_cv)
abline(v = log(ridge1_cv$lambda.min))
## 21 x 1 sparse Matrix of class "dgCMatrix"
## s1
## (Intercept) 0.132825360
## V1 1.334864503
## V2 0.031514349
## V3 0.739492669
## V4 0.048358071
## V5 -0.881162824
## V6 0.611469081
## V7 0.123949760
## V8 0.386769953
## V9 -0.038636714
## V10 0.118120656
## V11 0.245469489
## V12 -0.067686451
## V13 -0.045130346
## V14 -1.121405223
## V15 -0.130593351
## V16 -0.042130893
## V17 -0.045664317
## V18 0.060068396
## V19 -0.003373719
## V20 -1.108136513
## Perform adaptive LASSO with 10-fold CV
alasso1_cv <- cv.glmnet(x = x_cont, y = y_cont,
type.measure = "mse",
nfold = 10,
alpha = 1,
penalty.factor = 1 / abs(best_ridge_coef),
keep = TRUE)
## Penalty vs CV MSE plot
plot(alasso1_cv)
## 21 x 1 sparse Matrix of class "dgCMatrix"
## s1
## (Intercept) 0.1252790
## V1 1.3872769
## V2 .
## V3 0.7552762
## V4 .
## V5 -0.8925767
## V6 0.5683429
## V7 .
## V8 0.3623266
## V9 .
## V10 .
## V11 0.1748004
## V12 .
## V13 .
## V14 -1.1146180
## V15 .
## V16 .
## V17 .
## V18 .
## V19 .
## V20 -1.1239692
library(VGAM)
plot(y = dlaplace(seq(-3,3, length.out = 1000), scale = 1),
x = seq(-3,3, length.out = 1000), type = "l",
main = "Laplace distribution", ylab = "density", xlab = "")
## stan_glm
## family: gaussian [identity]
## formula: y ~ .
## observations: 100
## predictors: 21
## ------
## Median MAD_SD
## (Intercept) 0.1 0.1
## x.1 1.4 0.1
## x.2 0.0 0.1
## x.3 0.8 0.1
## x.4 0.1 0.1
## x.5 -0.9 0.1
## x.6 0.6 0.1
## x.7 0.1 0.1
## x.8 0.4 0.1
## x.9 0.0 0.1
## x.10 0.1 0.1
## x.11 0.3 0.1
## x.12 -0.1 0.1
## x.13 -0.1 0.1
## x.14 -1.2 0.1
## x.15 -0.1 0.1
## x.16 -0.1 0.1
## x.17 -0.1 0.1
## x.18 0.1 0.1
## x.19 0.0 0.1
## x.20 -1.1 0.1
##
## Auxiliary parameter(s):
## Median MAD_SD
## sigma 1.0 0.1
##
## ------
## * For help interpreting the printed output see ?print.stanreg
## * For info on the priors used see ?prior_summary.stanreg
## (Intercept) x.1 x.2 x.3 x.4 x.5
## 0.110978758 1.383406675 0.023433696 0.767465639 0.066199186 -0.903181023
## x.6 x.7 x.8 x.9 x.10 x.11
## 0.619143984 0.125173599 0.402498695 -0.038407595 0.134344297 0.252355538
## x.12 x.13 x.14 x.15 x.16 x.17
## -0.070953063 -0.050532662 -1.165130179 -0.145895451 -0.054105129 -0.056000871
## x.18 x.19 x.20
## 0.050886785 -0.004531585 -1.144454727
## stan_glm
## family: gaussian [identity]
## formula: y ~ .
## observations: 100
## predictors: 21
## ------
## Median MAD_SD
## (Intercept) 0.7 0.3
## x.1 0.0 0.0
## x.2 0.0 0.0
## x.3 0.0 0.0
## x.4 0.0 0.0
## x.5 0.0 0.0
## x.6 0.0 0.0
## x.7 0.0 0.0
## x.8 0.0 0.0
## x.9 0.0 0.0
## x.10 0.0 0.0
## x.11 0.0 0.0
## x.12 0.0 0.0
## x.13 0.0 0.0
## x.14 0.0 0.0
## x.15 0.0 0.0
## x.16 0.0 0.0
## x.17 0.0 0.0
## x.18 0.0 0.0
## x.19 0.0 0.0
## x.20 0.0 0.0
##
## Auxiliary parameter(s):
## Median MAD_SD
## sigma 2.9 0.2
##
## ------
## * For help interpreting the printed output see ?print.stanreg
## * For info on the priors used see ?prior_summary.stanreg
## (Intercept) x.1 x.2 x.3 x.4
## 6.525370e-01 2.349271e-03 5.342229e-04 9.873049e-04 -8.862767e-06
## x.5 x.6 x.7 x.8 x.9
## -1.045235e-03 1.267102e-03 1.813045e-04 5.819743e-04 -1.306971e-04
## x.10 x.11 x.12 x.13 x.14
## -3.313507e-04 5.830675e-04 -2.901755e-04 2.406221e-04 -2.050376e-03
## x.15 x.16 x.17 x.18 x.19
## 1.978188e-04 1.548878e-04 2.683929e-04 1.558121e-04 -1.629086e-05
## x.20
## -1.090514e-03
post_means <-
QuickStartExample %>%
rename(scale = y) %>%
filter(x.1 > 100)
for (i in 1:100){
laplace_scale <- seq(0.1, 0.001, length.out = 100)[i]
blm_laplace <- stan_glm(y ~ ., data = QuickStartExample, seed = 1,
prior = laplace(location = 0, scale = laplace_scale),
iter = 1000,
refresh = 0)
post_means <- rbind(post_means, c(coef(blm_laplace)[-1], laplace_scale))
}
library(reshape2)
names(post_means) <- c(names(QuickStartExample)[-21], "scale")
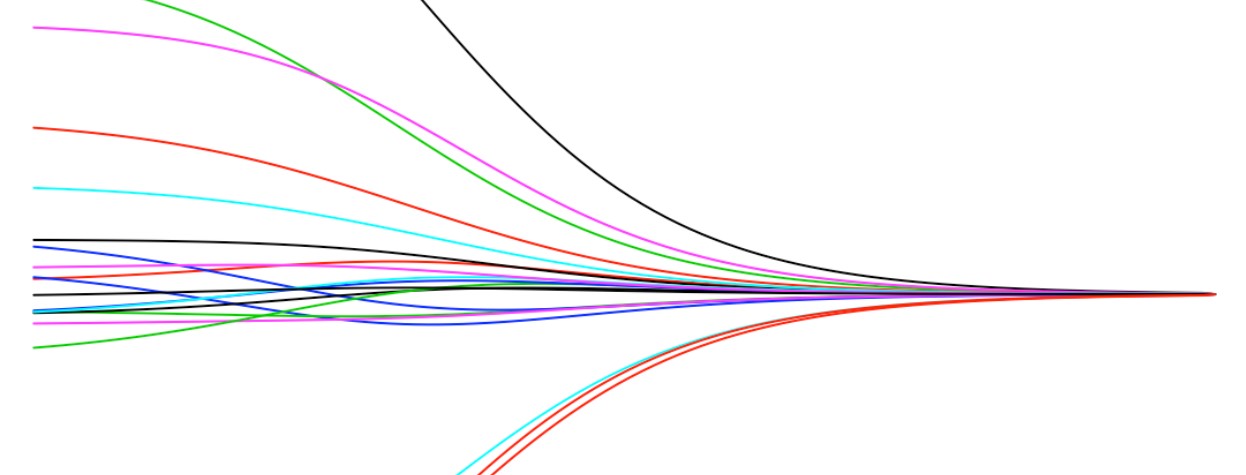
post_means %>%
melt(id.vars = "scale") %>%
mutate(scale_inv = 1/scale) %>%
ggplot(aes(x = scale_inv, y = value, col = variable)) +
scale_x_continuous(trans='log10') +
geom_line() +
theme_bw()